3’ end additions by T7 RNA polymerase are RNA self-templated, distributive, and diverse in character – RNA-Seq analyses
Yasaman Gholamalipour, Aruni Karunanayake Mudiyanselage, & Craig T. Martin, Nucl Acids Res, 46, 9253-9263, 2018.
Selected by the editors as a “Breakthrough Article.”
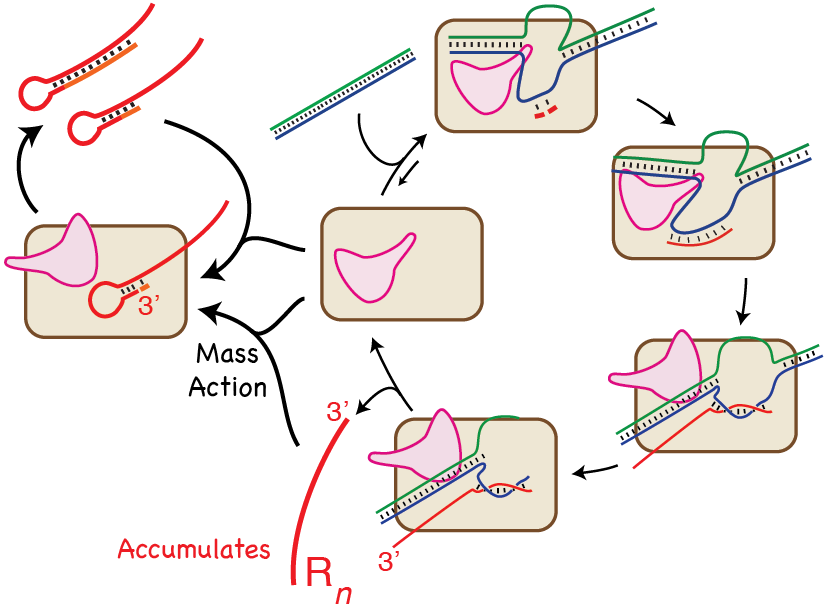
Synthetic RNA is widely used in basic science, nanotechnology and therapeutics research. The vast majority of this RNA is synthesized in vitro by T7 RNA polymerase or one of its close family members. However, the desired RNA is generally contaminated with products longer and shorter than the DNA-encoded product. To better understand these undesired byproducts and the processes that generate them, we analyze in vitro transcription reactions using RNA-Seq as a tool. The results unambiguously confirm that product RNA rebinds to the polymerase and self-primes (in cis) generation of a hairpin duplex, a process that favorably competes with promoter driven synthesis under high yield reaction conditions. While certain priming modes can be favored, the process is heterogeneous, both in initial priming and in the extent of priming, and already extended products can rebind for further extension, in a distributive process. Furthermore, addition of one or a few nucleotides, previously termed ‘nontemplated addition,’ also occurs via templated primer extension. At last, this work demonstrates the utility of RNA-Seq as a tool for in vitro mechanistic studies, providing information far beyond that provided by traditional gel electrophoresis.
Practical tip: never allow product RNA to accumulate to high concentrations (or watch this web site for more soon...)
PMID: 30219859 PMCID: 6182178 DOI: 10.1093/nar/gky796