Research
RNA is seeing explosive growth both as a therapeutic and as a target of therapeutics. In the COVID-19 era, mRNA vaccines have rightly shot to prominence, but mRNA therapeutics for disease treatment are right behind. CRISPR-based therapeutics, using inactivated Cas9, offer a wide range of therapeutic options, as well. These technologies all require long RNA (100-5,000 bases or more) and in vitro transcription using T7 RNA polymerase is either the preferred or the only option for manufacturing specific RNA sequences.
However, in vitro transcription, particularly when pushed to high production yields, is plagued by unwanted side reactions. We have recently characterized these interactions and have demonstrated that the products are very heterogeneous in nature, yet they all arise from self-primed extension of the target RNA. This results in RNA that is at least partially doubled stranded, and it is well-known that double stranded RNAs trigger our innate immune response - leading to the failure of RNA therapeutics in clinical trials.
Our goal for the near future is to develop approaches that eliminate these products and improve, more generally, the synthesis of RNA in high yield and purity. Towards this end, we exploit our mechanistic understanding of this complex system.
The Martin Lab
The Martin Lab has a long history of studying fundamental mechanisms in transcription (see our structure-based mechanistic summary!). Using powerful tools of biophysical chemistry and enzymology, we have focused on the (relatively) simple, single subunit RNA polymerase from bacteriophage T7. Our work is all with purified enzyme and largely with synthetic DNA templates, affording us exquisite control over the system.
Using tools from enzymology and biophysical chemistry, we have probed initial promoter binding, de novo initiation, initial transcription, the transition to elongation, and the elongation complex itself. Incorporation of fluorescent base analogs in the DNA provide direct read out of the polymerase as it moves through these phases. Detailed kinetics, combining fluorescence and more traditional polymerase assays, has proven invaluable in our studies. More generally, the ability to use synthetic DNA, with a wide variety of subtle or not so subtle base or backbone modifications has provide us with very detailed hypothesis testing. To learn more about our results, see the "About Transcription" tab above.
New directions complement existing tools
A few years back, a laboratory project put us, for the first time, in the place of "end-users" of T7 RNA polymerase. In trying to synthesize RNA at high yield, we encountered problems that many before us have encountered over the years. In particular, the polymerase was making the correct product, but was also making a variety of unintended/undesired RNA products.
Gel electrophoresis. Using gel electrophoresis, the entire polymerase(s) field analyzes products based on length. If we are expecting a product 40 bases long, but see one 35 bases in length, we can simply delete the last 5 bases from our intended sequence to "discern" the nature of the 35mer (or so we all think).
Longer than expected products. Often in transcription, one sees gel products migrating at longer lengths than expected from the DNA (40 bases in the simple study below). Despite numerous earlier studies (Cazenave & Uhlenbeck, Triana-Alonso et al., Nacheva & Bernal-Herranz) clearly indicating that these products arise from an RdRp templated process, the field has generally called at least the shorter additions "nontemplated," suggesting a promiscuous terminal transferase activity. But the nature and amounts of these products vary greatly with the encoded RNA sequence (not just the terminal bases). Even single nucleotide additions are templated.
Shorter than expected products. These RNAs can arise from termination at specific sequences in the DNA (see below). They might also arise from RNA polymerase "falling off" (in a non-sequence dependent manner) prematurely (this is extremely rare, although the expected length polydispersity would suggest that they will be hard to detect). Shorter dsRNA products can arise from short abortive products binding to RNA, within the polymerase active site, similarly to the mechanism above, and priming extension.
Very long unexpected products. Some have suggested that the RNA polymerase "turns around" and begins using the nontemplate strand as a template. More likely, this arises from polymerase initiating synthesis from one end of duplex DNA, in a promoter-independent manner (Mu et al.). In any case, this is also sequence dependent and is driven by high concentrations of Mg(II).

RNA-Seq. The advent of deep sequencing, in particular RNA-Seq, presents an opportunity to complement gel electrophoresis in very powerful ways. Instead of assuming RNA sequence from (often approximate) size estimates from a gel, we can sequence the entire RNA. And more importantly, it gives us a pool of up to a million individual RNA sequences. At its simplest, we can bin these together to reproduce a gel - how many 30mers, how many 31mers? This yields precise length resolution and dynamic range (we can detect RNAs that are barely or not detectable on gels).
Of course, RNA-Seq gives us the RNA lengths, but it also directly gives us the sequence. Or more correctly, as noted above, it gives us a large number of individual sequences. We can count individual sequences and rank them by relative abundance. From that analysis, it becomes clear that there are multiple sequence classes. The added sequence, beyond that expected from runoff noted above, gives us important information on the source of the product - how did it arise?

"Nontemplated additions" are templated. The results of our initial study show unambiguously that additions beyond the expected RNA length are indeed templated by the RNA, as suggested by some earlier studies. We were even able to show unambiguously that this arises from self-templating: the products do not arise from two different RNAs annealing to prime extension in trans, but rather a single RNA molecule folds back on itself (in cis) to allow extension. The priming hairpins are heterogeneous: different folding structures lead to initial extension starting at different positions along the RNA (evidenced in RNA-Seq by different initial sequences following the encoded runoff sequence). Extension can also be distributive: a primer-extended RNA can release, form an alternative structure, and then rebind a different polymerase for a second round of extension (evidenced in RNA-Seq by different sequence "blocks" in the final sequence). We see that n+1 and n+2 products (also templated, but that's another story) get chased to longer products with time.
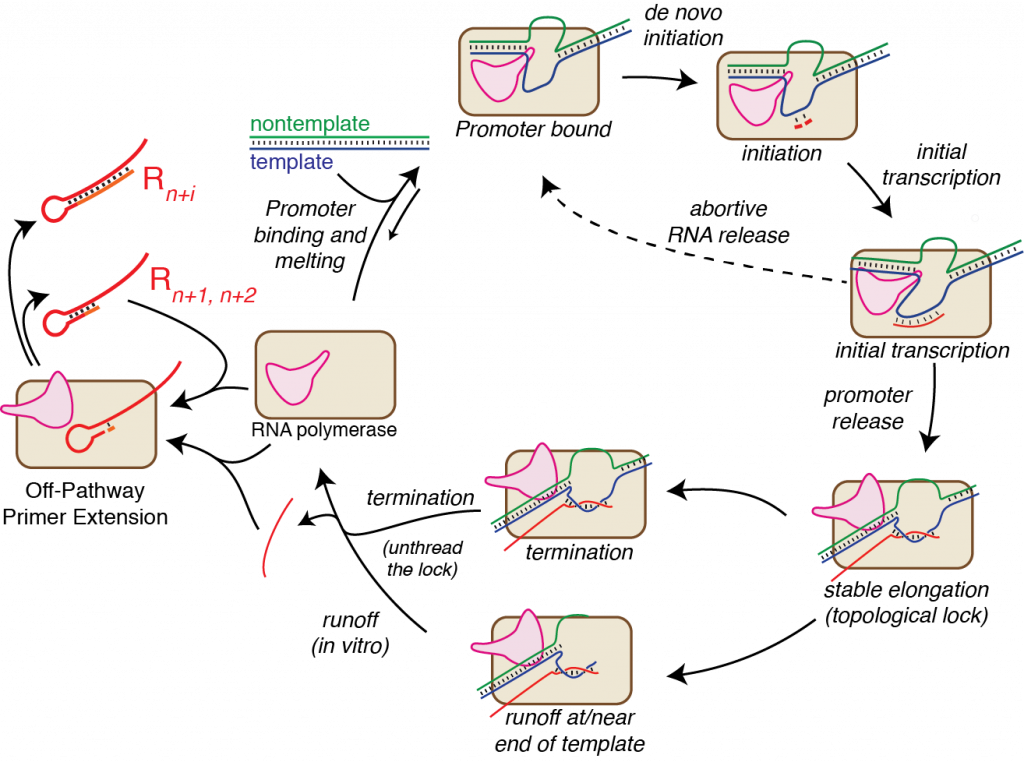
Self-primed extension is worse in preparative reactions. Since product RNA rebinds RNA polymerase, the principle of mass action predicts that this process will increase with increasing concentration of free RNA. Thus, initially in a reaction, the concentration of free RNA is small, and self-primed extension is minimal. But most users of T7 RNA polymerase aim for high yield of their RNA (we want a lot of RNA!). Sadly, mass action dictates that higher yield reactions will produce more self-primed extensions, increasing as the reaction proceeds.
Mechanistic understanding leads to solutions. Given new tools and new understandings, we are now embarking on a series of studies aimed at developing approaches toward RNA synthesis without the above "off-pathway" reactions. This will have dramatic impact in two ways: 1) since correct length (runoff) RNA will not get chased to longer lengths, the absolute yield of correct length RNA will increase and 2) the elimination of extended products from the final reaction mix may make labor intensive and low yield purification less necessary (and we're also working on better ways to do that!). Note that as RNA length increases, purification approaches are increasingly ineffective (preparatively separating a 200mer from a 205mer by gel or chromatographic approaches is simply not feasible).
But what about other bad things that polymerase does? RNA-Seq also tells us, as others have discerned before, that in addition to the above 3' heterogeneity, the final products also show 5' heterogeneity. So the n+2 product on the gel could be an RNA with two extra bases at the 3' end, or mis-initiation/misp-priming might have yielded an RNA with two additional bases at the 5' end. Though less frequent, we do see this in our RNA-Seq data. We are working on ways to prevent this as well.
Another class of "impurity" often seen in gels is comprised of shorter than encoded RNAs: n-1, n-2, etc. We very clearly observe these as well. We know that an elongation complex should become less stable as it approaches the end of a template (we have previously shown that this arises because barriers to forward translocation decrease near the end of a duplex template, not because of lost downstream duplex interactions). Solving this problem is more vexing, but we're working on it.
One of our goals is to develop a set of sequence guidelines: which sequences/structures yield the most problems, which the least? If you have an RNA sequence that has been problematic, please contact us. Maybe your sequence will confirm our findings or maybe, more interestingly(!), they'll challenge us. If your sequence is proprietary in some way, just send us the last (3'-most) 25-50 bases. We hypothesize that most of the trouble lies there.