Insights into the Mechanism of Initial Transcription in E coli RNA Polymerase. Energetic Mechanisms Leading to Initial Complex Instability and Abortive Cycling
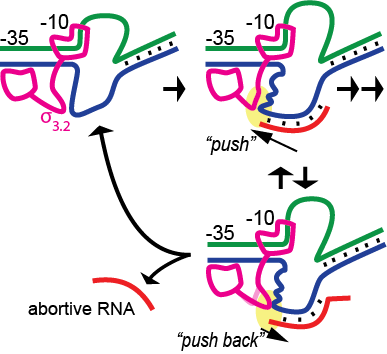
Satamita Samanta & Craig T. Martin, J. Biol. Chem. 288, 31993-32003, 2013.
It has long been known that during initial transcription of the first 8-10 bases of RNA, complexes are relatively unstable, leading to the release of short abortive RNA transcripts. An early stressed intermediate model led to a more specific mechanistic model proposing “scrunching” stress as the basis for the instability. Recent studies in the single subunit T7 RNA polymerase have argued against scrunching as the energetic driving force and instead argue for a model in which pushing of the RNA-DNA hybrid against a protein element associated with promoter binding, while likely driving promoter release, reciprocally leads to instability of the hybrid. In the current study, we test these models in the structurally unrelated multi-subunit bacterial RNA polymerase. Via the targeted introduction of mismatches and nicks in the DNA, we demonstrate that neither downstream bubble collapse nor compaction/scrunching of either the single stranded template or nontemplate strands is a major force driving abortive instability (although collapse from the downstream end of the bubble does contribute significantly to instability of artificially halted complexes). In contrast, pushing of the hybrid against a mobile protein element (sigma3.2 in the bacterial enzyme) results in substantially increased abortive instability and is likely the primary energetic contributor to abortive cycling. The results suggest that abortive instability is a byproduct of the mechanistic need to couple the energy of nucleotide addition (RNA chain growth) to driving the timed release of promoter contacts during initial transcription.
PMID: 24047893 PMCID: PMC3814795 DOI: 10.1074/jbc.M113.497669