Why better RNA?
The Martin lab, having studied fundamental mechanisms in transcription, is now on a quest to dramatically improve RNA synthesis. For almost four decades, the preferred approach for synthesizing RNA has been in vitro transcription by T7 RNA polymerase. We now understand, however, that current protocols are at times fundamentally flawed, producing impurities: RNAs other than directly encoded by the DNA. Some of these have double stranded regions, which can trigger the innate immune response, but RNA impurities of any kind can interfere with applications (and are often difficult to purify away).
Based on recent results from our lab (and from those before us), we are developing synthetic approaches that will keep the RNA impurities (byproducts of the reaction) from being synthesized in the first place. This will lead not only to higher purity from the outset, but also to higher yields.
Emerging technologies require synthetic RNA
The following briefly describes just some of the emerging technologies that require long (>50 base) RNA in high purity.
mRNA Therapeutics. Genetic diseases generally result from a defect (mutation) in the DNA that encodes RNA, which (often, but not always) encodes a now mutant protein. This can result in "loss of function," with debilitating consequences. Protein replacement therapies aim to provide a patient with "good" (native) versions of the protein, to restore function. This requires synthesizing the protein in a lab and then formulating the protein for delivery to the patient. Although there are some astounding successes, this process my be re-developed for every new protein (disease).
mRNA therapeutics aims to deliver a "good" (native) copy of the mRNA, so that the patient's own cells can then produce the "good" (native) protein. In principle, this has many advantages over direct protein replacement. Different protein products are simply different sequences of RNA, which can all be formulated the same. By producing the proteins in patient, the proteins receive correct post-translational modifications.
However, early attempts at mRNA-driven protein replacement led to an unintended stimulation of the immune system (inflammation), and in some cases, this was fatal. We believe that unintended byproducts of the in vitro production of mRNA are responsible for this immune response. Indeed, it has been shown that highly purified mRNA can avoid the immune response.
mRNA Vaccines. One traditional approach to vaccines is to administer to patients inactivated (or fragmented) protein that is unique to the invading organism. The patient's body then generates antibodies that can serve to rapidly fight off future threats.
An alternative approach, related to mRNA therapeutics, is to deliver to patients mRNA encoding the protein or protein fragment. The patient's cells then generate the protein, which then elicites antibody production.
In this case, while some stimulation of the immune system is desired, one would prefer to have control over that stimulation and not rely on impurities to do the job.
CRISPR Technologies. There are a wide variety of emerging technologies that exploit the novel CRISPR system. Central to the unique targeting ability of CRISPR is "guide RNA," which is delivered therapeutically as a part of the CRISPR system. As above, immune-stimulating RNA impurities can lead to (sometimes dangerous) side effects.
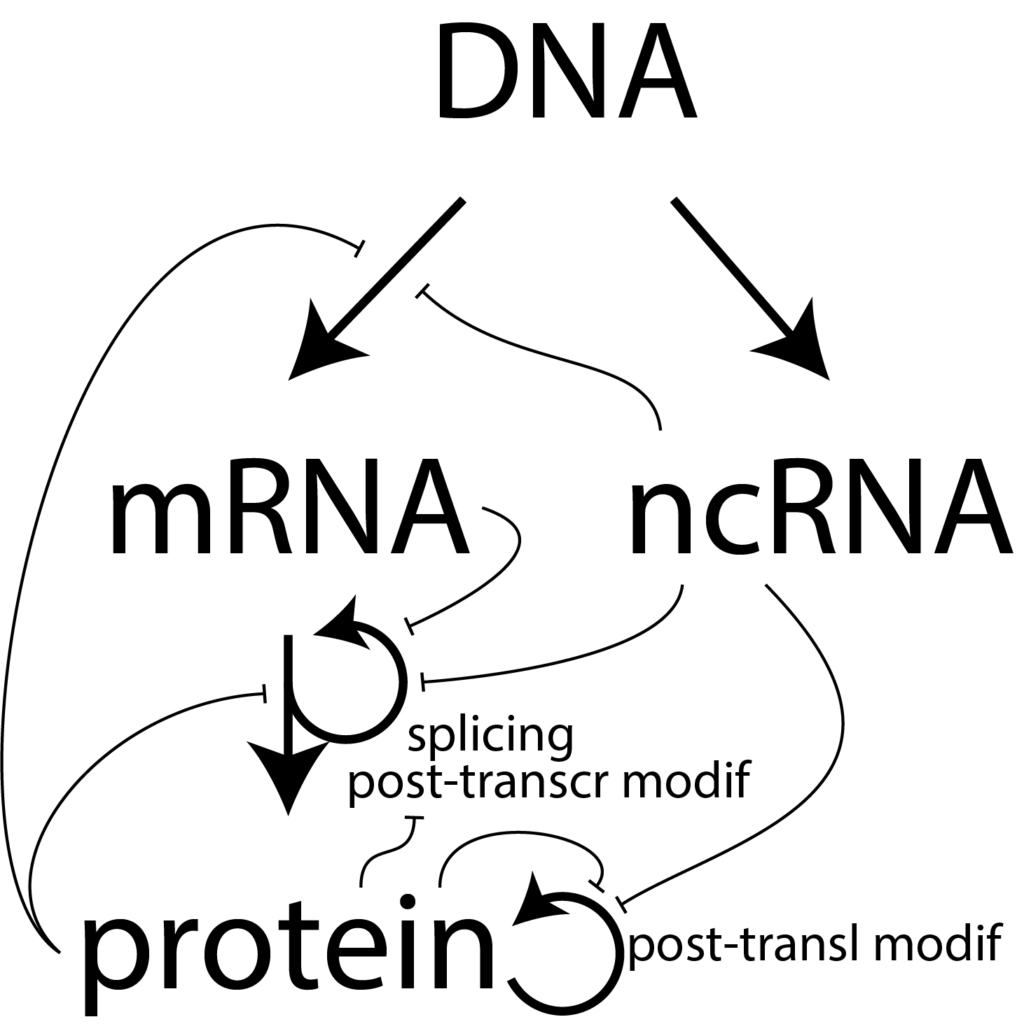
Cellular RNA as a Drug Target. We used to think that RNA's only/primary function was encoding protein (and that is important!). Discoveries of the last few decades, however, have pointed to a wide variety of roles for "noncoding" RNA in human biology. Alternative splicing of RNA is increasingly appreciated for the complex role it plays in regulating cells. Indeed, the instructions for splicing often reside within the noncoding (spliced out) portions of mRNA.
We also now appreciate that (non-mRNA) long noncoding (ncRNA) RNAs themselves play key roles in a wide variety of processes, including neural function. In this case, genetic diseases can arise from improper splicing, alterations in post-transcriptional modifications, or changes in RNA stability in the cell.
In addition to designer RNA therapeutics to treat such diseases, traditional (small molecule) therapeutics that bind to specific (generally structured) RNA have the potential for huge impact on health. Screens of such drugs, however, require pure in vitro RNA targets - RNA free of RNA-based impurities.
Emerging RNA technologies
The complexity of cellular RNA in cells is now appreciated across a wide variety of organisms. From diagnostics, to crop management, to