Unit Activity of T7 RNA polymerase
The definition of unit activity in T7 RNA polymerase is an outdated terminology that is not used uniformly. The general definition is the amount of nucleotide incorporated per unit time.
In theory, one can use the equation below to convert to the more appropriate measure of molar concentration. The molar mass of T7 RNA polymerase is 98,855 g/mol. The specific activity is often reported as ≈400,000 U/mL, but different vendors use different values, varying by as much as a factor of 3.
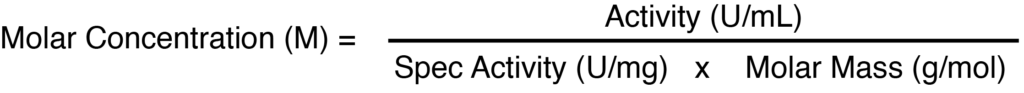
The original definition was based on using the 40 kb phage genomic (with 17 promoters) DNA as template. For practical reasons, modern definitions use different DNA templates, but different vendors use different DNA templates. Different vendors also use different reaction conditions.
A sampling of vendor definitions follows:
New England Biolabs - 1 Unit is defined as the amount of enzyme that will incorporate 1 nmol ATP into acid-insoluble material in a total reaction volume of 50 μl in 1 hour at 37°C. Components: 40 mM Tris-HCl (pH 7.9 @ 25°C), 1 mM DTT, 2 mM spermidine, 6 mM MgCl2, 0.5 mM each ATP, UTP, GTP, CTP, and 20 µg/mL T7 DNA.
Promega- 1 Unit is defined as the amount of enzyme required to catalyze the incorporation of 5 nmol of CTP into acid-insoluble product in 60 minutes at 37°C in a total volume of 100 μl. Components: 40mM Tris-HCl (pH 7.9 at 25°C), 6 mM MgCl2,10 mM DTT, 10 mM NaCl, 2 mM spermidine, 0.05% Tween® 20, 0.5 mM each of ATP, GTP, CTP, and UTP, 0.5 μCi [3H]CTP and 20 µg/mL supercoiled pGEM®-5Zf(+) Vector DNA.
Sigma-Aldrich - 1 Unit is defined as the amount of enzyme that synthesizes 1 μg of mRNA per hour in a 30-minute reaction containing 25 μg/mL linearized plasmid, DNA 3 mM of each NTP and 18 mM MgCl2, at 37 °C, pH 7.7.
Original Definition - 1 Unit is defined as the amount of enzyme which (in a 10 min reaction in 100 µL) gives a rate of 1 nmole of AMP incorporation in 1 h at 37° C, in 40 mM Tris pH 8, 20 mM MgCl2, 0.4 mM each NTP, 0.01 mM BME, 1 mg/mL BSA, 360 µM (phage genomic) DNA*
*The original paper (M Chamberlin, McGrath, & Waskell, 1970) notes 0.036 µmol phage DNA in 100 µL. If this is 0.036 µmol bases, then the molar concentration of 40 kb DNA would be 9 nM (with 17 promoters, ≈150 µM in promoter). This seems most likely.
Note that what matters for promoter binding/initiation is molar concentration (of enzyme and of DNA). Only the original 1970,paper provides that. Citing milligrams of template DNA is not useful, unless we know the length of the DNA, to allow conversion to a molar concentration. Thus, even though all three vendors include 20-25 µg/mL template DNA, the molar concentration of promoter can be different in each case. Length of the transcript also matters - for a 1 kb transcript, initiation is mostly rate limiting, but for a 10 kb transcript, elongation is mostly rate limiting (for the same molar concentration of template).